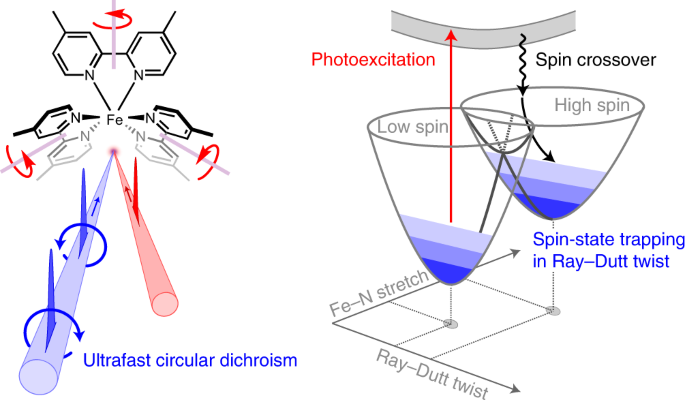
Mechanism of rotenone binding to respiratory complex I depends on ligand flexibility | Scientific Reports

The binding mode of orphan glycyl-tRNA synthetase with tRNA supports the synthetase classification and reveals large domain movements | Science Advances
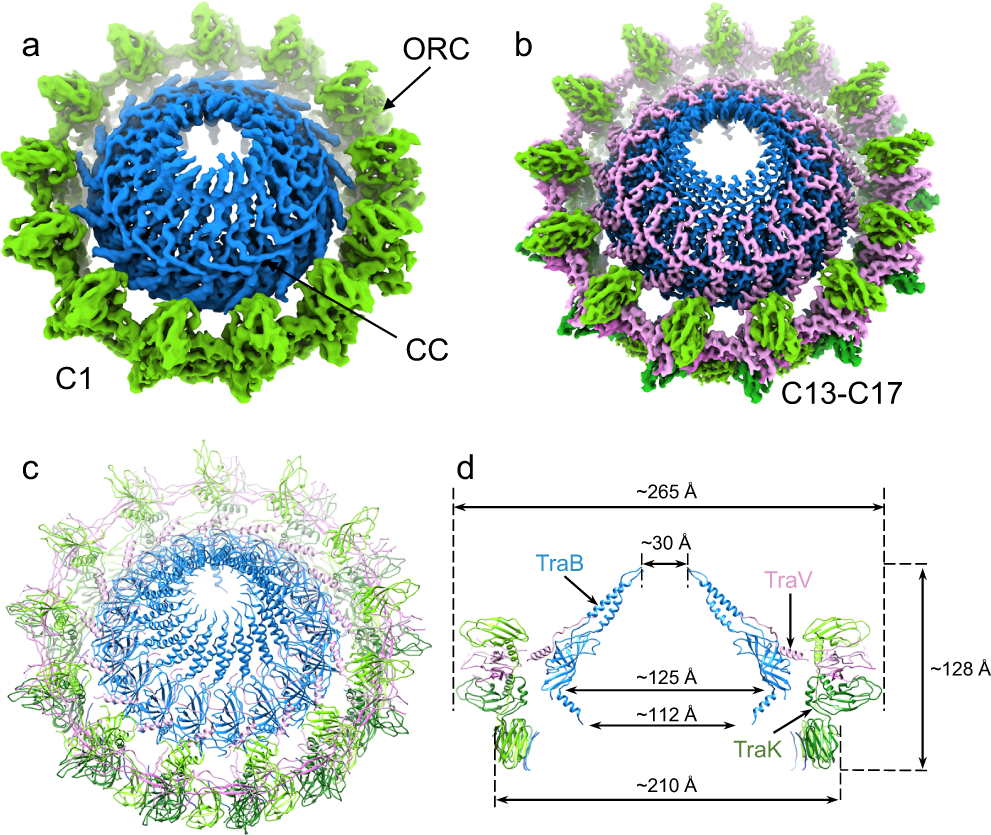
Structure of a type IV secretion system core complex encoded by multi-drug resistance F plasmids | Nature Communications

A trivalent 4f complex with two bis-silylamide ligands displaying slow magnetic relaxation | Nature Chemistry

Photochemical Identification of Auxiliary Severe Acute Respiratory Syndrome Coronavirus 2 Host Entry Factors Using μMap | Journal of the American Chemical Society
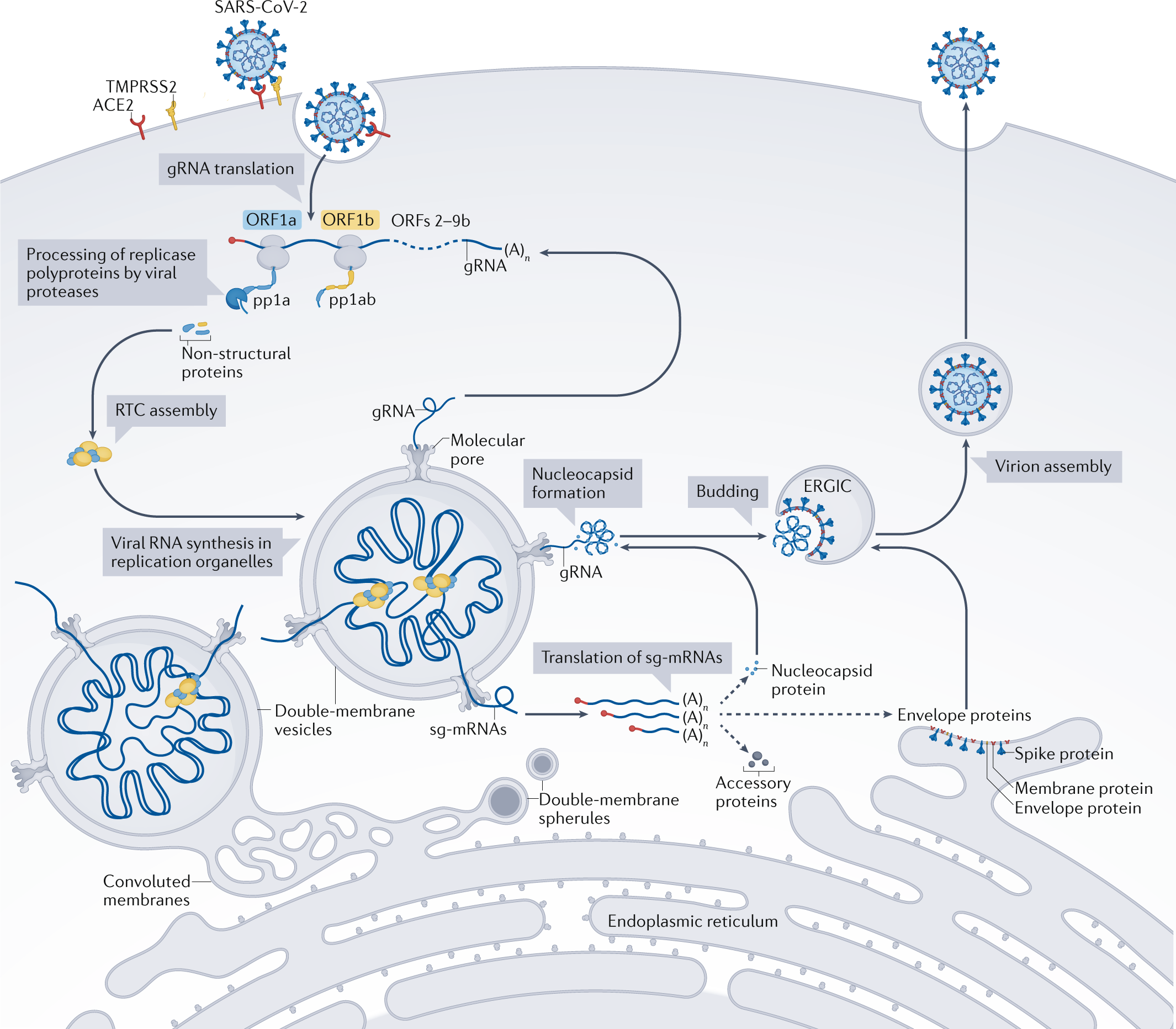
Structures and functions of coronavirus replication–transcription complexes and their relevance for SARS-CoV-2 drug design | Nature Reviews Molecular Cell Biology

High-Throughput Analysis Reveals miRNA Upregulating α-2,6-Sialic Acid through Direct miRNA–mRNA Interactions | ACS Central Science

Potent neutralization of SARS-CoV-2 variants of concern by an antibody with an uncommon genetic signature and structural mode of spike recognition - ScienceDirect
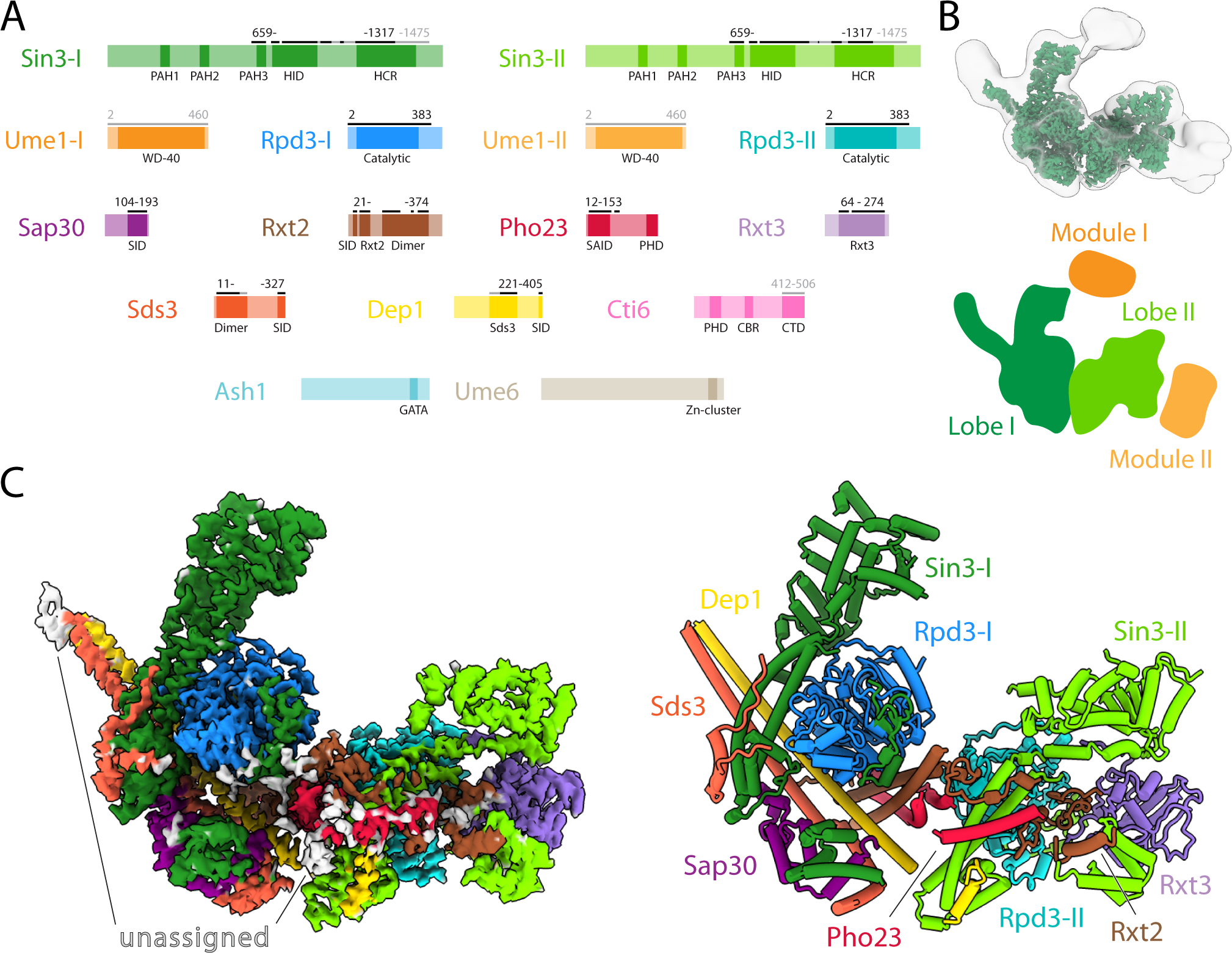
Cryo-EM structure of the Saccharomyces cerevisiae Rpd3L histone deacetylase complex | Nature Communications
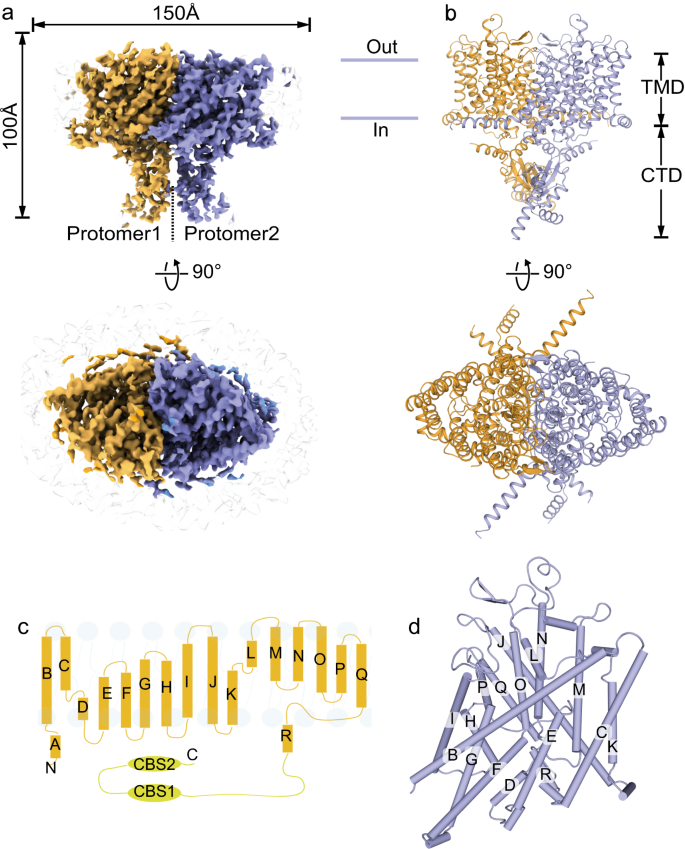
Cryo-EM structures of ClC-2 chloride channel reveal the blocking mechanism of its specific inhibitor AK-42 | Nature Communications

Exploration of Type II Binding Mode: A Privileged Approach for Kinase Inhibitor Focused Drug Discovery? | ACS Chemical Biology
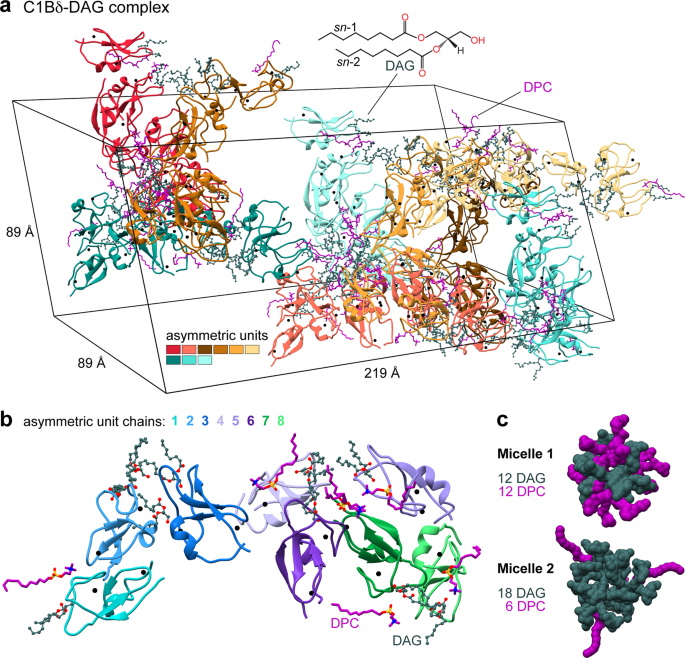
Structural anatomy of Protein Kinase C C1 domain interactions with diacylglycerol and other agonists | Nature Communications