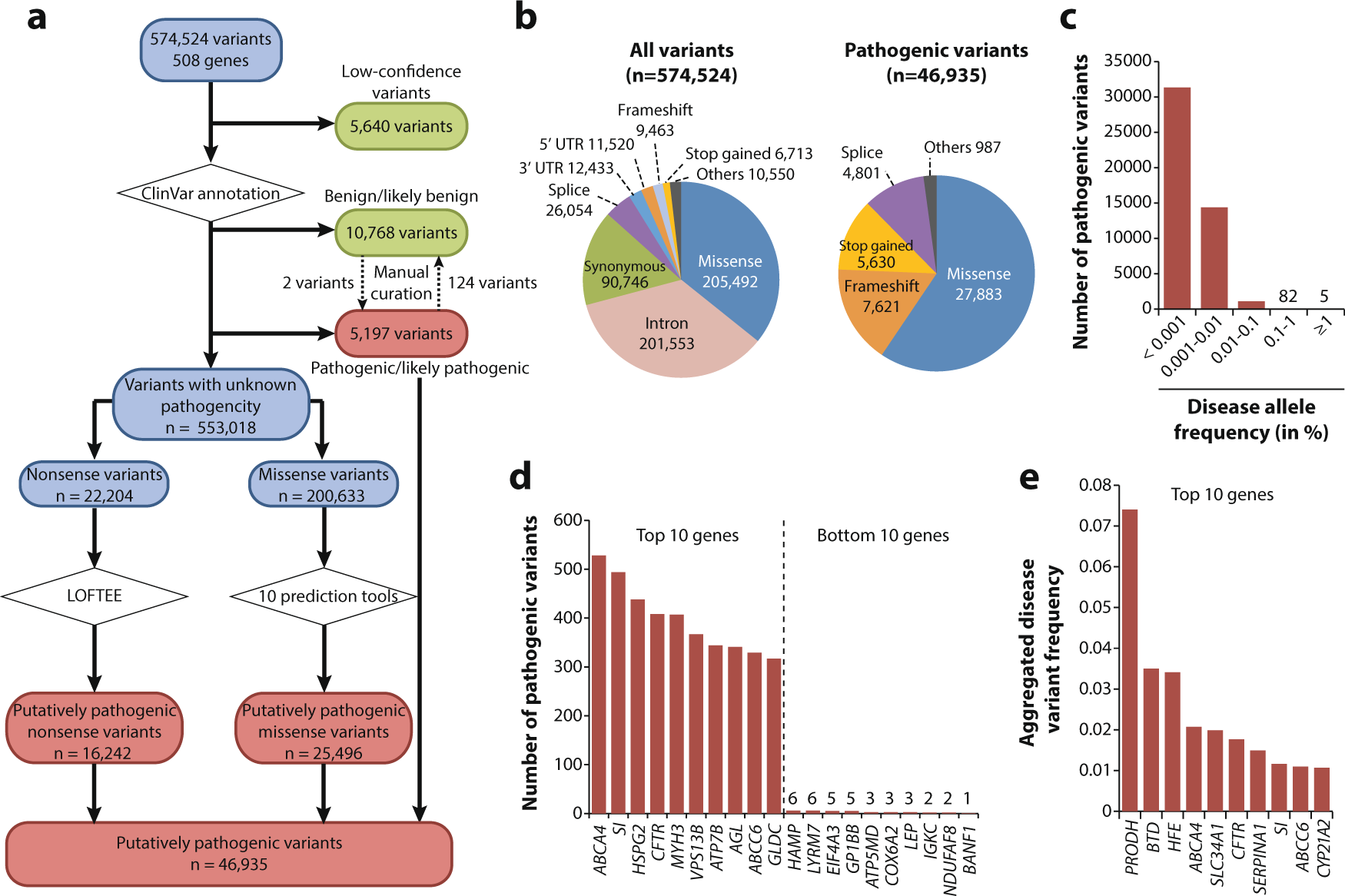
The prevalence, genetic complexity and population-specific founder effects of human autosomal recessive disorders | npj Genomic Medicine
Estimating Additive and Non-Additive Genetic Variances and Predicting Genetic Merits Using Genome-Wide Dense Single Nucleotide Polymorphism Markers | PLOS ONE

Estimation of individual animal SNP-BLUP reliability using full Monte Carlo sampling - ScienceDirect

Translating polygenic risk scores for clinical use by estimating the confidence bounds of risk prediction | Nature Communications

The reliability and heritability of cortical folds and their genetic correlations across hemispheres | Communications Biology

Patterns of Reliability: Assessing the Reproducibility and Integrity of DNA Methylation Measurement - ScienceDirect
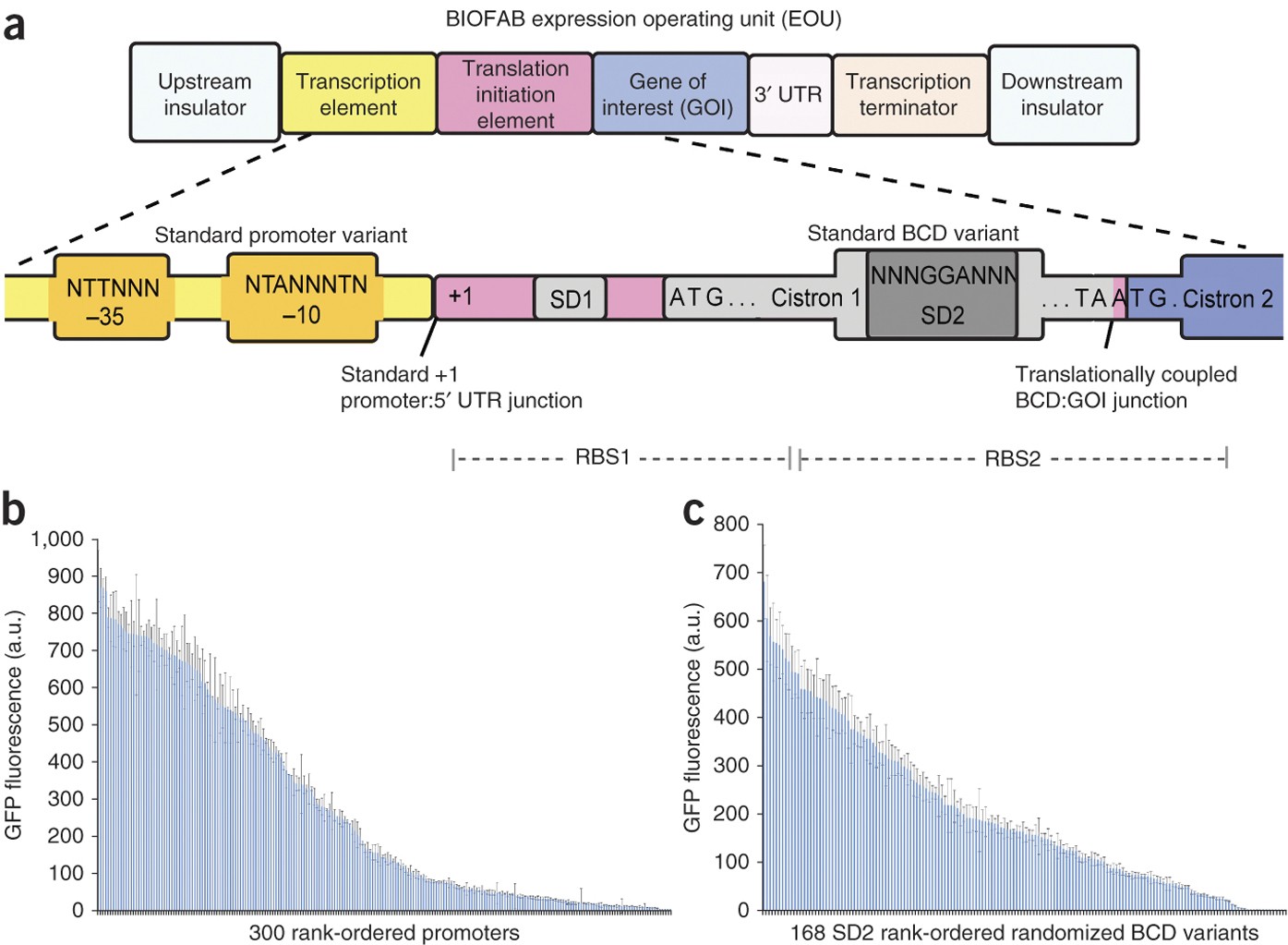
Precise and reliable gene expression via standard transcription and translation initiation elements | Nature Methods
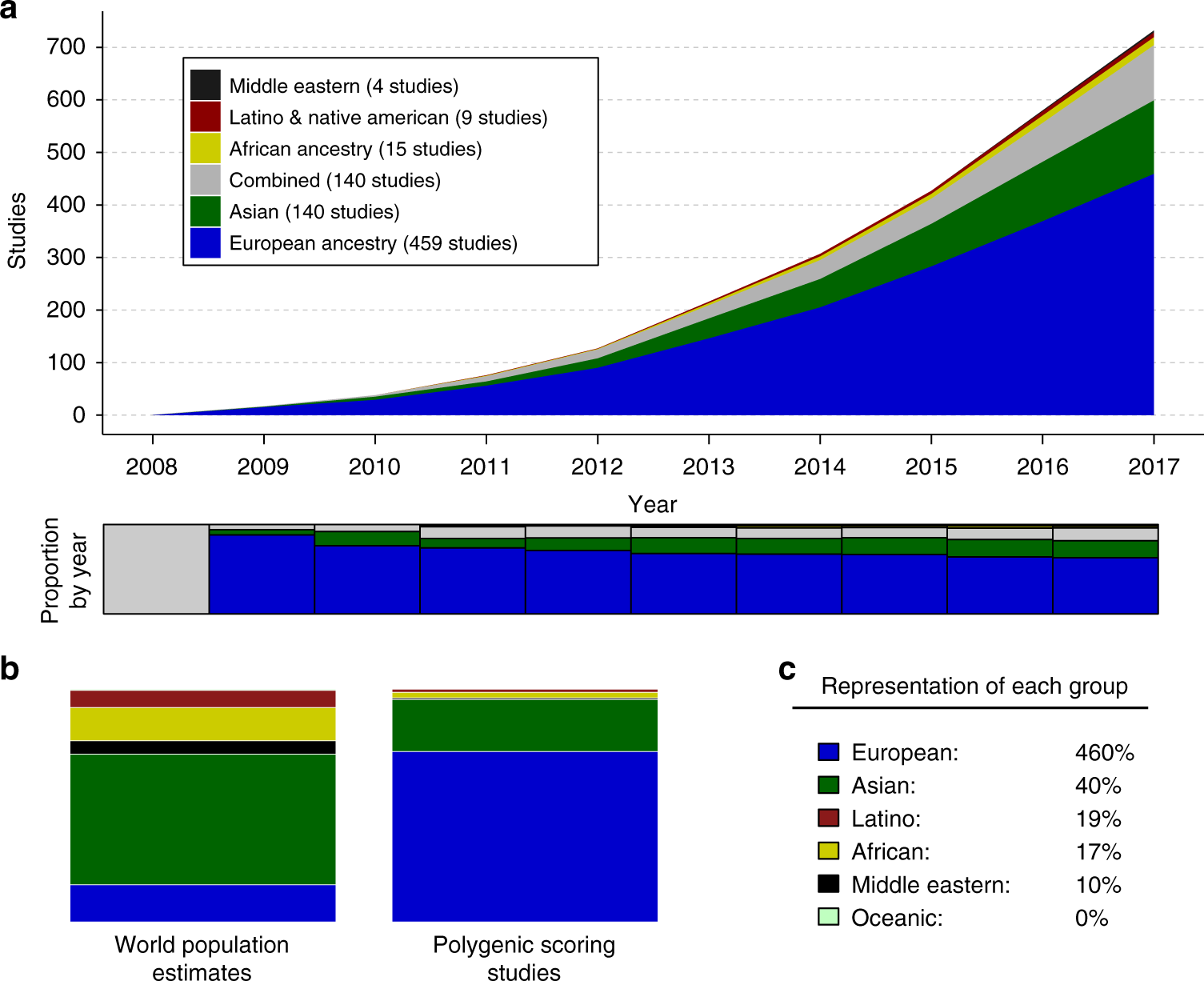
Analysis of polygenic risk score usage and performance in diverse human populations | Nature Communications
Test-retest reliability of the HEXACO-100—And the value of multiple measurements for assessing reliability | PLOS ONE
Survival Analysis in Python (KM Estimate, Cox-PH and AFT Model) | by Rahul Raoniar | The Researchers' Guide | Medium
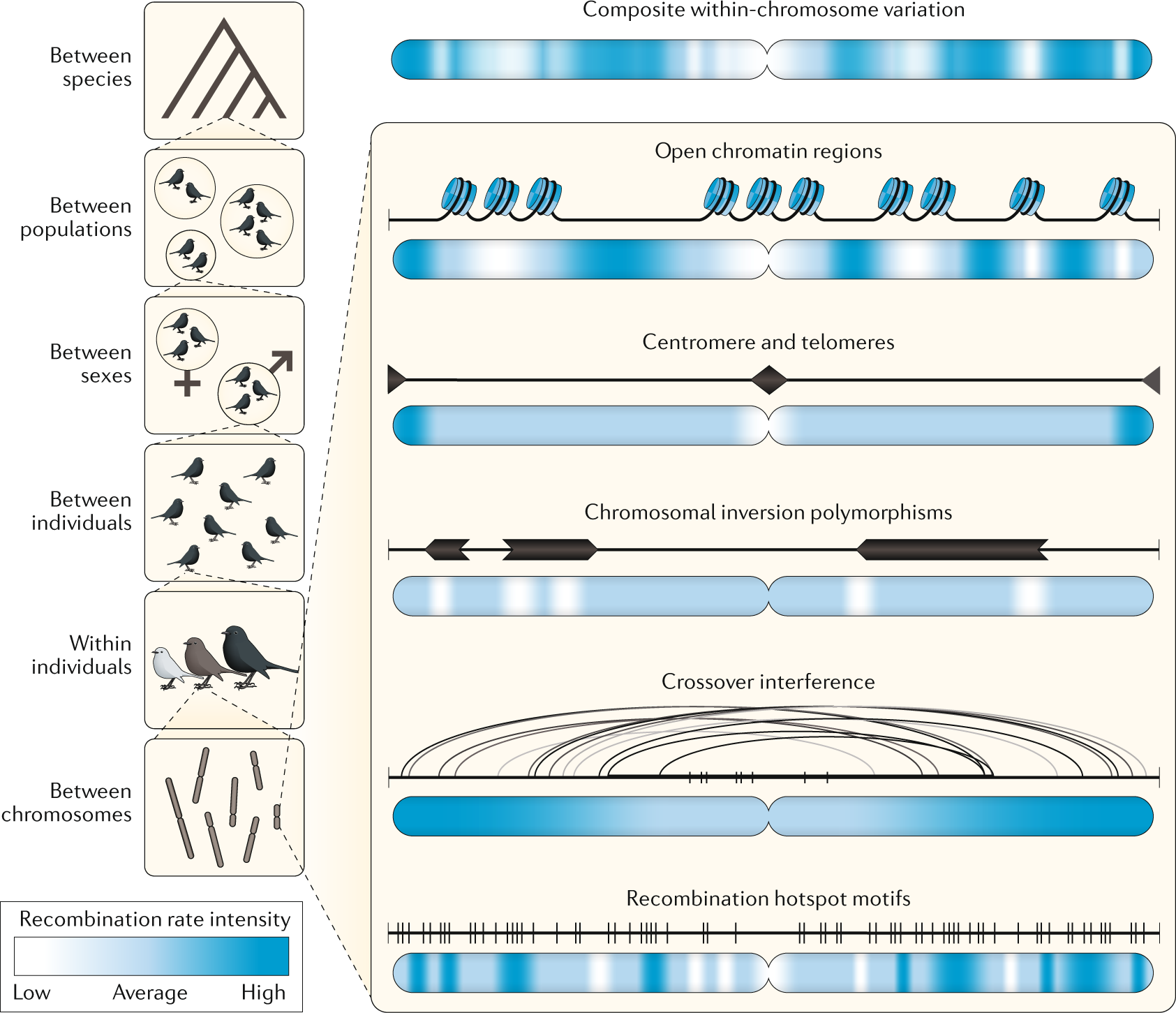
From molecules to populations: appreciating and estimating recombination rate variation | Nature Reviews Genetics

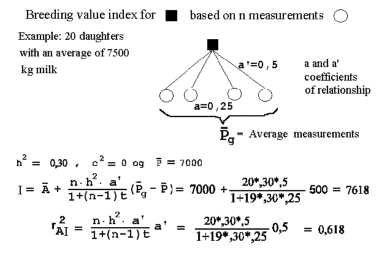
Appendix D: methods for obtaining approximate reliability for genetic evaluations. | Linear models for the prediction of animal breeding values